Machine Learning
| Title | Description | Input Data | |
|---|---|---|---|
MLflow Example Model (Wine Preference) | This is an example Capsule to show the functionality of the MLFLow Orchestrator Capsule. It generates a model, logs parameters and results into the mlflow database and then syncs the results to S3. |
| |
MLflow Orchestrator Capsule | This is a central MLflow "Orchestrator" Capsule that can be used to interactively view the results from multiple machine learning Capsules using the same MLflow database |
| |
MLFlow Example Model (Diabetes Experiment) | This is an example Capsule to show the functionality of the MLFLow Orchestrator Capsule. It generates a model, logs parameters and results into the mlflow database and then syncs the results to S3. |
| |
DECIMER 2.0 | This Capsule uses DECIMER (Deep Earning for Chemical ImagE Recognition) 2.0 to predict the SMILES (simplified molecular input line entry system) for input chemical images. |
| |
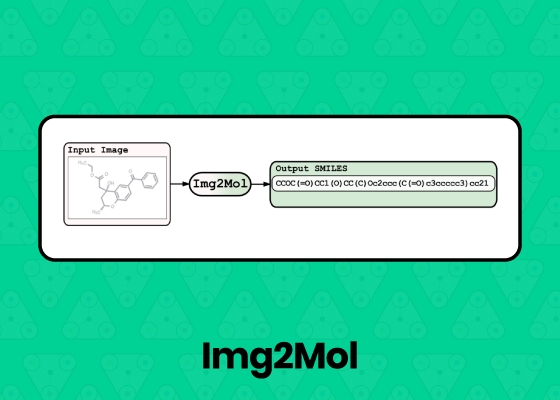
Img2Mol | This Capsule uses Img2Mol to predict the compound structure for input chemical images. It will output a structure data file and SMILES representation for each input image. |
| |
Streamlit ColabFold: AlphaFold2 using MMseqs2 | ColabFold offers an accelerated prediction of protein structures and complexes by combining the fast homology search of MMseqs2 with AlphaFold2. This Capsule can be accessed through a Streamlit Cloud Workstation, run in a Pipeline or accessed using the App Panel. |
| |
CombFold - prepare | This Capsule generates fasta files of protein subunits for downstream processing by AlphaFold. The subunits are then assembled using Streamlit ColabFold: AlphaFold2 using MMseqs2 |
| |
CombFold- Combinatorial Assembly | The CombFold Combinatorial Assembly algorithm assembles subunits from AlphaFold into a single large complex. |
|